Blog: Deconstructing the Heterogeneity of Tumors via Single-Cell Sequencing
Key takeaways:
- Cancerous tumors have complex and heterogeneous characteristics, and the variations within and between tumors influence disease progression and treatment.
- Recent advances in single-cell RNA sequencing allow researchers to analyze individual cells and subpopulations within a tumor, revealing intricate details that are masked during bulk sequencing.
- Huge strides in machine learning and statistical analysis are enabling sophisticated data insights and increasing understanding of tumor heterogeneity.
- The ability to tailor cancer treatment based on the specific characteristics of a patient's tumor is a significant advancement, enabling more targeted therapies and better patient outcomes.
- Cancer research is beset with challenges such as ethical dilemmas, high costs, and the availability of technologies, but single-cell sequencing and its potential to revolutionize cancer care offer a promising future.
Cancer has a deep historical footprint, which has been proven by archeologists unearthing fossilized bone tumors in human mummies from ancient Egypt, and references to tumors found in ancient manuscripts. Intra-tumor heterogeneity, a well-established phenomenon in human cancers, highlights the intricacies of this disease. Modern research reveals that cancerous tumors are not solitary entities but rather heterogeneous assemblies of cells, each possessing a unique genetic profile. This diversity significantly influences the disease's progression, patients' resistance to treatment, and the ultimate prognosis. Single-cell RNA sequencing offers the potential to deconstruct intra-tumor heterogeneity and personalize cancer treatments for better outcomes.
Understanding Intra-Tumor Heterogeneity
Tumor heterogeneity has significant implications for cancer research and treatment. With intra-tumor heterogeneity, variations exist within a single tumor, including different cell types or regions. These differences are influenced by unique genetic mutations and environmental factors like oxygen levels, diet, exposure to harmful substances, and even previous treatments.
This complexity makes diagnosing and treating cancer more challenging, as different tumor parts may respond differently to therapy, and a sample from one area might not represent the whole tumor. Additionally, tumors can change and adapt, adding further difficulty to predicting their behavior and treating them effectively. Understanding this multifaceted nature of tumors is vital for personalized medicine and requires sophisticated tools and knowledge.
Comprehending Genetic Diversity
Comprehending how genetic diversity evolves over time remains a formidable challenge. Historically, a major hurdle in deciphering the dynamics of tumor evolution was the difficulty in obtaining longitudinal samples from cancer patients. Consequently, most prior studies, relying heavily on bulk analysis from single time-point samples, could offer only indirect insights into tumor evolution.
However, the emergence of single-cell RNA sequencing technologies has transformed our capabilities. It allows us to explore tumor evolution with unparalleled precision by dissecting the genetic diversity within tumors at the individual cell level. This advancement revolutionizes our ability to comprehend and track the progression of the disease over time.
Single-Cell RNA Seq and the Role of Bioinformatics
Single-cell RNA sequencing provides invaluable insights into this complex tumor landscape. It empowers researchers to unravel the cellular complexity of tumors and paves the way for more personalized and effective therapeutic strategies. I firmly believe that single-cell sequencing has the potential to revolutionize cancer treatment concerning tumors, and to enable us to understand smaller, treatment-resistant populations within tumors with greater precision and effectiveness.
Yet, the true essence of single-cell sequencing extends beyond mere data generation. The real value emerges in the interpretation of this data. Bioinformatics plays a pivotal role in this aspect, transforming raw sequences into meaningful insights that can guide research and treatment. By analyzing these datasets, we can uncover patterns, anomalies, and unique characteristics that might otherwise remain hidden in the vast sea of genetic information.
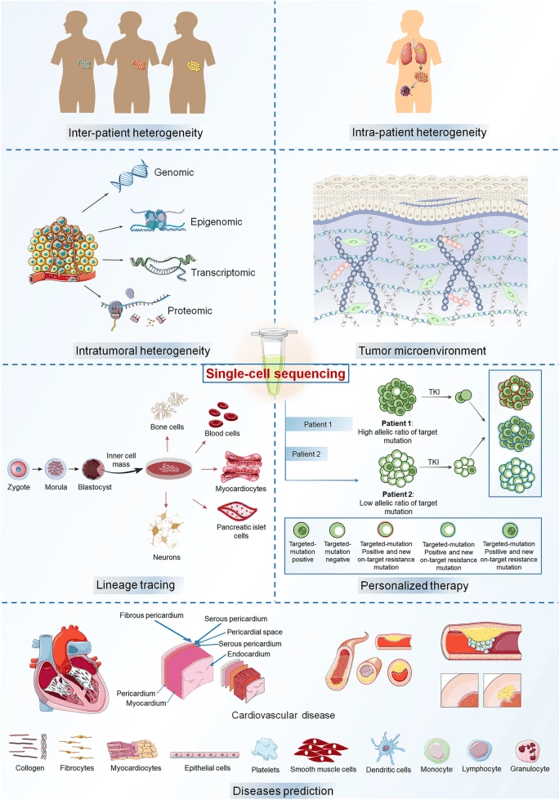
|
Illustrations of single-cell RNA sequencing applications in different fields. (1) tumor heterogeneity, (2) tumor microenvironment, (3) lineage tracing, (4) personalized therapy, and (5) disease prediction Image, caption source: Chang, X., Zheng, Y. & Xu, K. Single-Cell RNA Sequencing: Technological Progress and Biomedical Application in Cancer Research. Mol Biotechnol (2023). Creative Commons license |
Revolutionizing Research with Single-Cell RNA Seq
Single-cell RNA sequencing represents a significant advancement over bulk sequencing, giving us unprecedented insight into the complex tumor landscape. Traditional sequencing methods analyze bulk tumor samples, averaging the genetic information of thousands or millions of cells. In contrast, single-cell sequencing allows researchers to analyze the genetic material of individual cells within a tumor.
The process enables us to identify the different subpopulations of tumors, including aggressive cancerous cells, immune cells, and other stromal cells, each with distinct genetic and phenotypic characteristics. This information is crucial for understanding the mechanisms driving tumor progression, resistance to therapy, and metastasis. It paves the way for more targeted and effective therapeutic strategies, supporting the shift toward personalized cancer treatment.
Making Single-Cell RNA Seq Accessible
Recent advancements in single-cell sequencing are making this technology more accurate and accessible for cancer research and treatment. Innovations have led to the development of more efficient cell isolation techniques, such as microfluidic platforms, which will enable the precise capture of individual cells. The introduction of new high-throughput sequencing platforms will increase the speed and reduce the cost of sequencing, making it feasible for broader applications.
Enhanced bioinformatics tools and computational algorithms will improve the accuracy of data analysis, allowing for more robust detection of genetic variations and expression patterns. Collaborative efforts and shared databases will also foster a more accessible research environment, enabling scientists worldwide to benefit from shared knowledge and resources. However, while the advancements in bioinformatics will offer immense potential, they could also bring forth certain challenges.
Challenges of Single-Cell RNA Seq
Handling vast datasets from single-cell sequencing can be computationally intensive, and ensuring the quality of this data will be paramount. Integrating data from heterogeneous sources will add another layer of complexity. Moreover, the field must grapple with the need for standardized bioinformatics protocols, to ensure analyses are reproducible across different studies and platforms. With various tools and algorithms available, standardizing processes will be crucial to ensure consistent and reliable results.
However, these challenges are not insurmountable. The key lies in interdisciplinary collaboration. By fostering strong partnerships between biologists, clinicians, and computational scientists, we will bridge the gaps in understanding and leverage the strengths of each discipline. Such collaborations will be pivotal in navigating the complexities of bioinformatics and ensuring that the insights derived are both accurate and actionable.
Personalized Cancer Treatment
The ability to tailor cancer treatment based on the specific characteristics of a patient's tumor will be a significant advancement in personalized cancer care. By analyzing these characteristics, oncologists will identify specific mutations, pathways, or receptors that may drive the tumor's growth and behavior.
This information will enable the selection of targeted therapies designed to interfere with the particular mechanisms underlying that specific tumor. Tailoring treatment this way will increase the likelihood of therapeutic success, minimize unnecessary side effects, and lead to better patient outcomes.
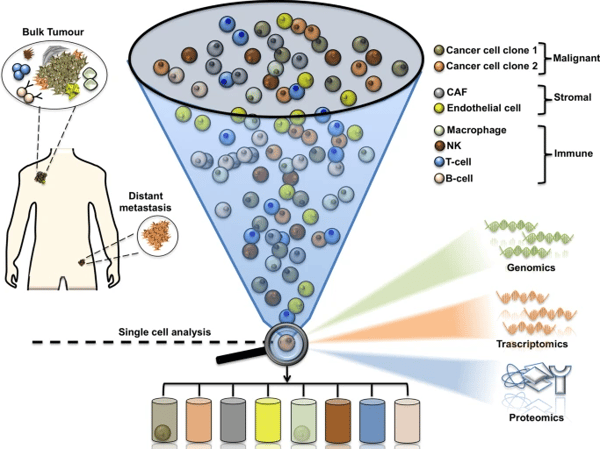
|
Schematic diagram illustrating single cell analysis ability to solve intratumor heterogeneity. Image, caption source: Fattore, L., Ruggiero, C.F., Liguoro, D. et al. Single cell analysis to dissect molecular heterogeneity and disease evolution in metastatic melanoma. Cell Death Dis 10, 827 (2019). https://doi.org/10.1038/s41419-019-2048-5. Creative Commons license: https://creativecommons.org/licenses/by/4.0/ |
Future Perspectives
The journey towards understanding and treating cancer is fraught with challenges and opportunities, including difficulties in sample preparation, sequencing, and data analysis; ethical dilemmas and regulatory compliance in research; and the high cost and limited availability of single-cell sequencing technologies.
Ongoing advancements in techniques, technologies, and computational methods will help to overcome these challenges. While single-cell RNA seq offers huge potential for cancer research and treatment, making the process robust and reproducible will be expensive and challenging.
New Horizons in Sequencing Technologies
The world of single-cell sequencing is quickly advancing with exciting new technologies and methods. Researchers are developing better ways to separate individual cells, making it easier to study them without causing damage. There are also new equipment and techniques that can read cells' genetic information faster, more accurately, and at a lower cost.
On the analysis side, we’re seeing new algorithmic techniques being applied to integrate the data types and then analyze that data. Scientists can now look at different layers of information within a single cell, giving a fuller picture of how it functions. In addition, we're starting to see techniques like single-cell ATAC-seq and other epigenetic approaches.
I believe we will soon start to see even more techniques that look into other features of the cells, like what proteins are expressed or what metabolites are found in each of these cells. Each piece of information gives more granularity or texture to the cell state.
Charting a New Course for Cancer Treatments
The journey through the complex landscape of tumor heterogeneity reveals a world where every tumor is a unique puzzle. Single-cell sequencing has emerged as a powerful tool, allowing scientists to dissect this complexity, cell by cell, and uncover the hidden variations that make each tumor distinct.
The advancements in technology, the move towards more accessible and accurate methods, and the integration of ethical considerations will all pave the way for a future where cancer care is more precise, effective, and tailored to the individual. While this technology is currently expensive and challenging, it will mature in due course. I believe in the next three to five years we’ll start seeing the impact of this research at the clinical level.
Read more about single cell RNA seq:
David AR, Zimmerman MR. Cancer: an old disease, a new disease or something in between? Nat Rev Cancer. 2010 Oct;10(10):728-33. doi: 10.1038/nrc2914. Epub 2010 Sep 3. PMID: 20814420.
Alizadeh AA, Aranda V, Bardelli A, Blanpain C, Bock C, Borowski C, Caldas C, Califano A, Doherty M, Elsner M, Esteller M, Fitzgerald R, Korbel JO, Lichter P, Mason CE, Navin N, Pe'er D, Polyak K, Roberts CW, Siu L, Snyder A, Stower H, Swanton C, Verhaak RG, Zenklusen JC, Zuber J, Zucman-Rossi J. Toward understanding and exploiting tumor heterogeneity. Nat Med. 2015 Aug;21(8):846-53. doi: 10.1038/nm.3915. PMID: 26248267; PMCID: PMC4785013.
Eirew P, Steif A, Khattra J, Ha G, Yap D, Farahani H, Gelmon K, Chia S, Mar C, Wan A, Laks E, Biele J, Shumansky K, Rosner J, McPherson A, Nielsen C, Roth AJ, Lefebvre C, Bashashati A, de Souza C, Siu C, Aniba R, Brimhall J, Oloumi A, Osako T, Bruna A, Sandoval JL, Algara T, Greenwood W, Leung K, Cheng H, Xue H, Wang Y, Lin D, Mungall AJ, Moore R, Zhao Y, Lorette J, Nguyen L, Huntsman D, Eaves CJ, Hansen C, Marra MA, Caldas C, Shah SP, Aparicio S. Dynamics of genomic clones in breast cancer patient xenografts at single-cell resolution. Nature. 2015 Feb 19;518(7539):422-6. doi: 10.1038/nature13952. Epub 2014 Nov 26. PMID: 25470049; PMCID: PMC4864027.
Heath JR, Ribas A, Mischel PS. Single-cell analysis tools for drug discovery and development. Nat Rev Drug Discov. 2016 Mar;15(3):204-16. doi: 10.1038/nrd.2015.16. Epub 2015 Dec 16. PMID: 26669673; PMCID: PMC4883669.
Haque A, Engel J, Teichmann SA, Lönnberg T. A practical guide to single-cell RNA-sequencing for biomedical research and clinical applications. Genome Med. 2017 Aug 18;9(1):75. doi: 10.1186/s13073-017-0467-4. PMID: 28821273; PMCID: PMC5561556.
Lähnemann D, Köster J, Szczurek E, McCarthy DJ, Hicks SC, Robinson MD, Vallejos CA, Campbell KR, Beerenwinkel N, Mahfouz A, Pinello L, Skums P, Stamatakis A, Attolini CS, Aparicio S, Baaijens J, Balvert M, Barbanson B, Cappuccio A, Corleone G, Dutilh BE, Florescu M, Guryev V, Holmer R, Jahn K, Lobo TJ, Keizer EM, Khatri I, Kielbasa SM, Korbel JO, Kozlov AM, Kuo TH, Lelieveldt BPF, Mandoiu II, Marioni JC, Marschall T, Mölder F, Niknejad A, Raczkowski L, Reinders M, Ridder J, Saliba AE, Somarakis A, Stegle O, Theis FJ, Yang H, Zelikovsky A, McHardy AC, Raphael BJ, Shah SP, Schönhuth A. Eleven grand challenges in single-cell data science. Genome Biol. 2020 Feb 7;21(1):31. doi: 10.1186/s13059-020-1926-6. PMID: 32033589; PMCID: PMC7007675.

Mark Kunitomi
Mark Kunitomi is the Chief Scientific Officer at Almaden Genomics. He was a post-doctoral fellow at UC San Francisco with a background in genomics, bioinformatics and microbiology, and he has a Ph.D. in Biochemistry & Molecular Biology from the University of California, San Francisco.